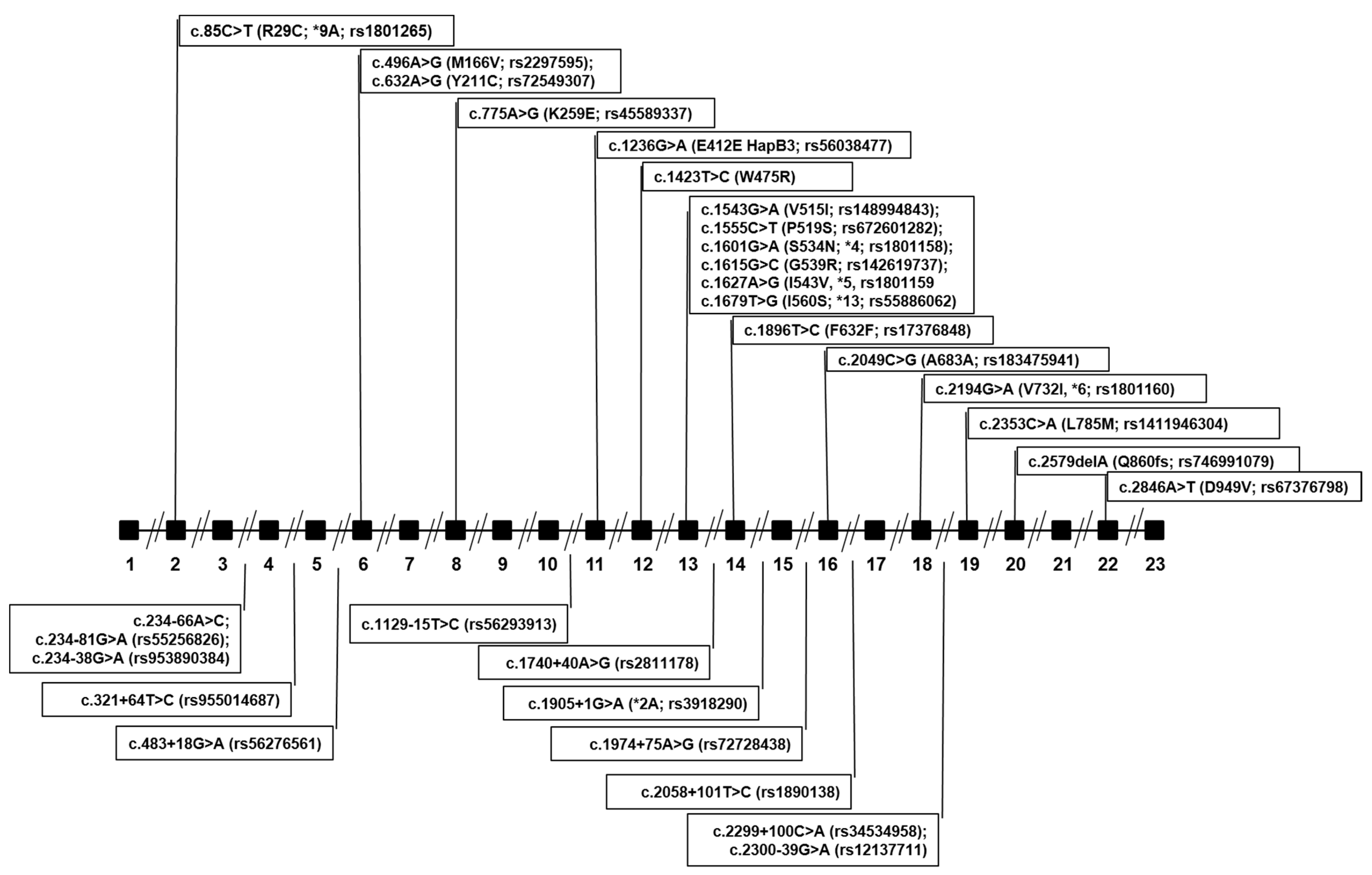
IJMS | Free Full-Text | Predicting Dihydropyrimidine Dehydrogenase Deficiency and Related 5-Fluorouracil Toxicity: Opportunities and Challenges of DPYD Exon Sequencing and the Role of Phenotyping Assays
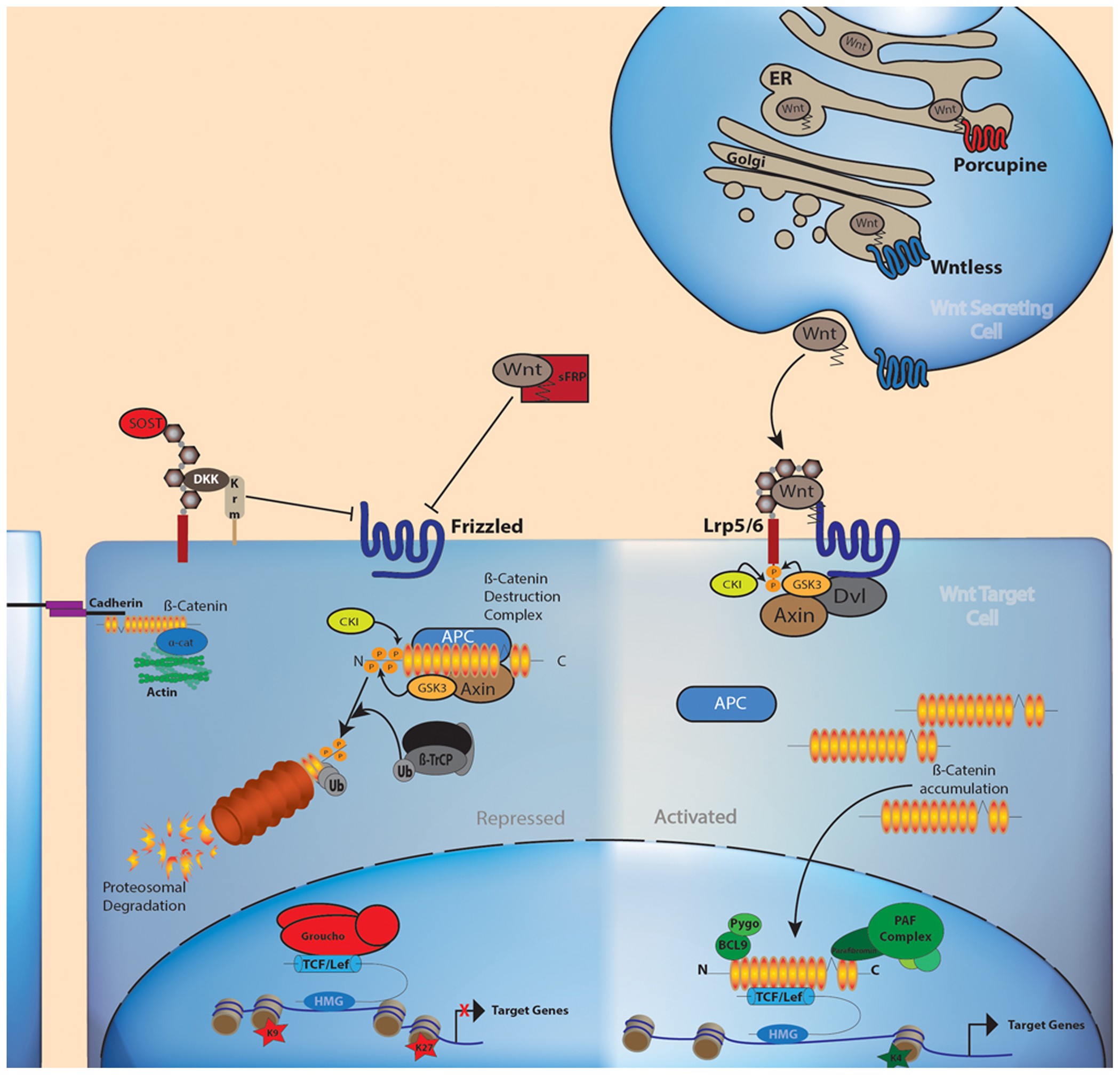
A Comprehensive Overview of Skeletal Phenotypes Associated with Alterations in Wnt/β-catenin Signaling in Humans and Mice | Bone Research
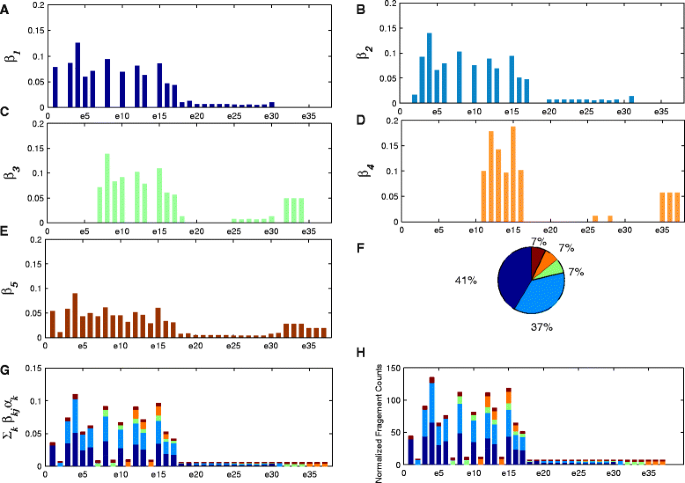
Improving RNA-Seq expression estimation by modeling isoform- and exon-specific read sequencing rate | BMC Bioinformatics | Full Text

PDF) The exon quantification pipeline (EQP): A comprehensive approach to the quantification of gene, exon and junction expression from RNA-seq data

Indazole-Based Covalent Inhibitors To Target Drug-Resistant Epidermal Growth Factor Receptor | Journal of Medicinal Chemistry
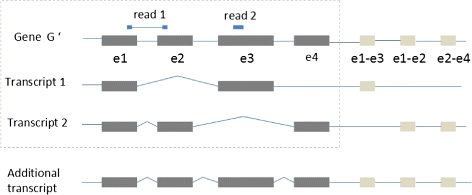
Improving RNA-Seq expression estimation by modeling isoform- and exon-specific read sequencing rate | BMC Bioinformatics | Full Text

Integrative (epi) Genomic Analysis to Predict Response to Androgen-Deprivation Therapy in Prostate Cancer - eBioMedicine

LongGF: computational algorithm and software tool for fast and accurate detection of gene fusions by long-read transcriptome sequencing | BMC Genomics | Full Text

Dynamic analyses of alternative polyadenylation from RNA-seq reveal a 3′-UTR landscape across seven tumour types | Nature Communications
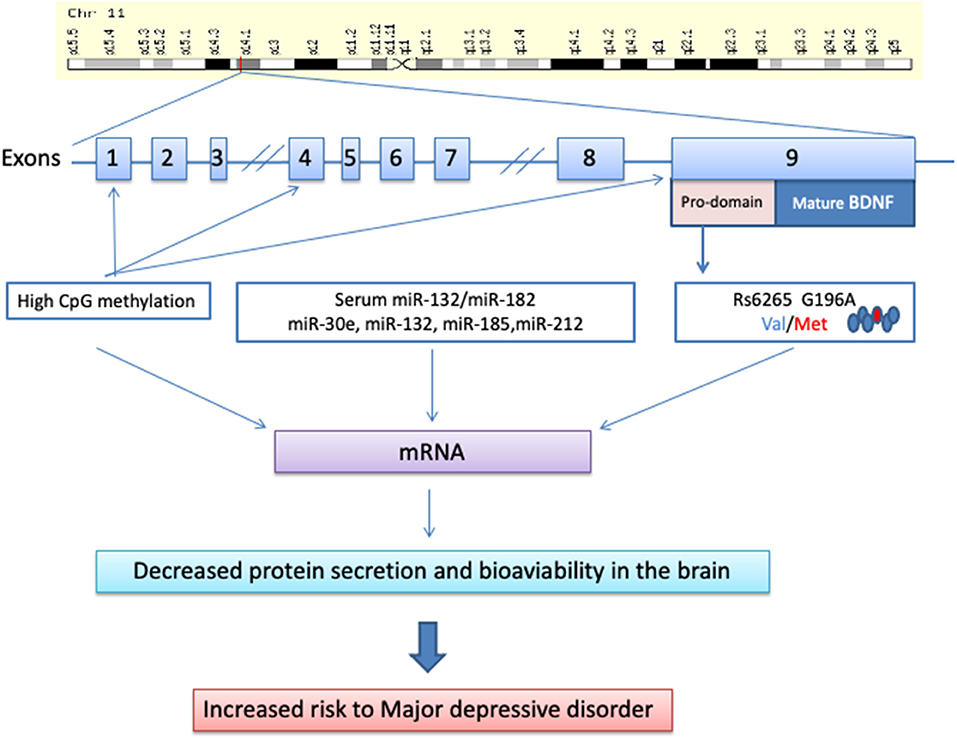
Frontiers | Blood Brain-Derived Neurotrophic Factor (BDNF) and Major Depression: Do We Have a Translational Perspective?